What is pairwise sequence alignment?
What is pairwise sequence alignment?
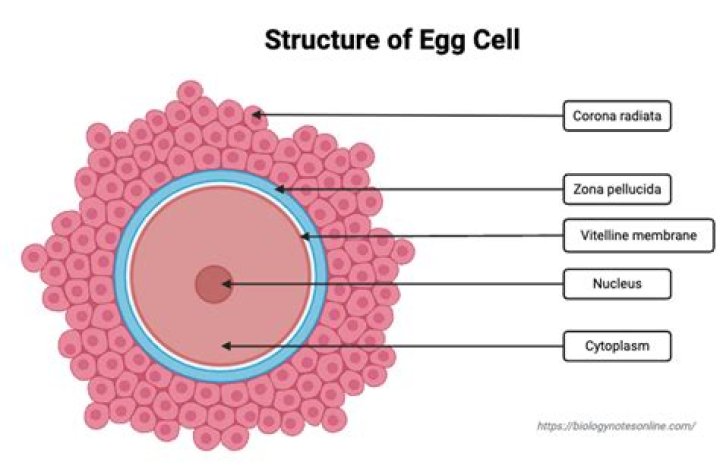
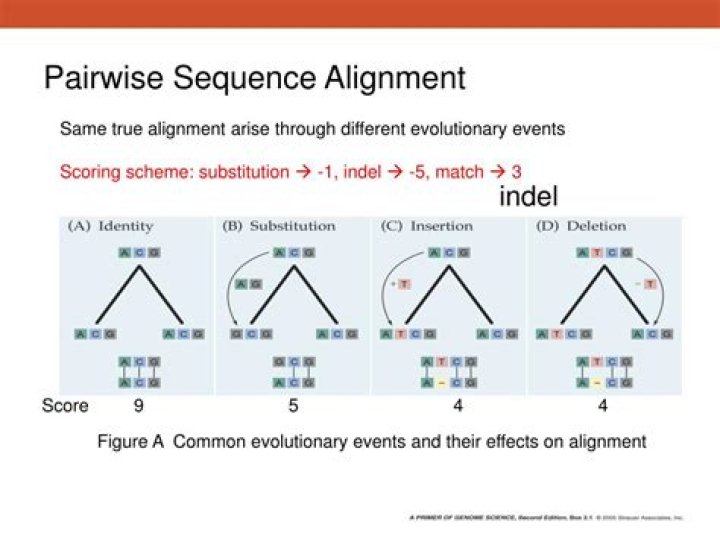
Pairwise Sequence Alignment is used to identify regions of similarity that may indicate functional, structural and/or evolutionary relationships between two biological sequences (protein or nucleic acid).
Which tools are used for pairwise sequence alignment?
Pairwise alignment
| Name | Description |
|---|---|
| LALIGN | Multiple, non-overlapping, local similarity (same algorithm as SIM) |
| NW-align | Standard Needleman-Wunsch dynamic programming algorithm |
| mAlign | modelling alignment; models the information content of the sequences |
| matcher | Waterman-Eggert local alignment (based on LALIGN) |
How is pairwise sequence alignment done?
In order to align a pair of sequences, a scoring system is required to score matches and mismatches. The scoring system can be as simple as “+1” for a match and “-1” for a mismatch between the pair of sequences at any given site of comparison.
What is emboss needle used for?
EMBOSS Needle reads two input sequences and writes their optimal global sequence alignment to file. It uses the Needleman-Wunsch alignment algorithm to find the optimum alignment (including gaps) of two sequences along their entire length.
What is pairwise identity?
Pairwise % Identity. When viewing alignments or assemblies this gives the average percent identity over the alignment. This is computed by looking at all pairs of bases at the same column and scoring a hit (one) when they are identical, divided by the total number of pairs.
What is pairwise score?
A Pairwise Algorithm is an algorithmic technique with its origins in Dynamic programming. Pairwise algorithms have several uses including comparing a protein profile (a residue scoring matrix for one or more aligned sequences) against the three translation frames of a DNA strand, allowing frameshifting.
Why do we use pairwise alignment?
What is ClustalW and what is its purpose?
ClustalW is a widely used system for aligning any number of homologous nucleotide or protein sequences. For multi-sequence alignments, ClustalW uses progressive alignment methods. The algorithm starts by computing a rough distance matrix between each pair of sequences based on pairwise sequence alignment scores.
What is the difference between embossed and engraved?
How Is Embossing Different? However, while an engraved style is achieved by removing trace amounts of material, embossing uses a die that raises the material – card, sheet metal, tin – to the same level from beneath, or varying levels using a multi-level die, to create a 3D impression.
What is geneious alignment?
Below is a brief overview of each algorithm. Geneious aligner. The Geneious aligner is a progressive pairwise aligner, similar to ClustalW (below). It is the slowest algorithm in Geneious and recommended for small alignments (e.g. fewer than 50 sequences, less than 1 kb in length). MUSCLE.
How is alignment score calculated?
The score of an alignment, S, calculated as the sum of substitution and gap scores. Substitution scores are given by a look-up table (see PAM, BLOSUM). Gap scores are typically calculated as the sum of G, the gap opening penalty and L, the gap extension penalty. For a gap of length n, the gap cost would be G+Ln.